IM Browser - a web application to query and visualize interaction networks
 |
Start IM BrowserRead the Quick Guide below first |
About IM Browser
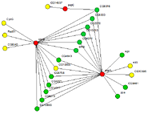
IM Browser runs as an applet inside of an internet browser. It provides a user-friendly interface to query interactome data. Users can search DroID for interactions involving one or many genes. Results are displayed as an interactive network graph where nodes represent genes/proteins and edges represent interactions. A network can be expanded, filtered, colored, and saved as an image or as a table of interactions and attributes. Queries can also be saved and rerun later to restore the graph.
Requirements
IM Browser works on Mac OS X with Safari. It may not work with Firefox or Chrome becuase of blocks to security settings for Java. IM Browser was implemented as a Java applet. To work with the applet you must have Java installed and enabled. IM Browser works well with Java version 6 and previous versions. Your Web Browser must allow java applets from http://www.droidb.org. The site can be entered in the Security settings of your your browser.
For Java 7
To use IM Browser with Java version 7 and later the Java security settings must allow applications to be launched from http://www.droidb.org. On a Mac go to System Preferences and Java. Under the Security tab edit the Exceptions List by adding http://www.droidb.org.
Quick Guide
After clicking "Start IM Browser" a new page opens and Java starts. If Java does not start successfully, check that you have the proper version and that Java is enabled in your web browser (see Requirements). When IM Browser loads successfully, open a Query Form using the menu "Add/Add Gene". Find the genes or proteins you are interested in (Step 1) and select the interaction data you wish to search (step 2), then click OK. The Query Form can be called up again by using the Add/Add Gene/Protein menu.
Tutorial
For a quick tutorial click here. It will take about 5 minutes to go through the tutorial.
Frequently Asked Questions
Lists some common problems and solutions.
Questions and feedback
Contact DroID@wayne.edu.